Note
Click here to download the full example code or to run this example in your browser via Binder
Running NiBetaSeries¶
This example runs through a basic call of NiBetaSeries using
the commandline entry point nibs.
While this example is using python, typically nibs will be
called directly on the commandline.
Import all the necessary packages¶
import tempfile # make a temporary directory for files
import os # interact with the filesystem
import urllib.request # grad data from internet
import tarfile # extract files from tar
from subprocess import Popen, PIPE, STDOUT # enable calling commandline
import matplotlib.pyplot as plt # manipulate figures
import seaborn as sns # display results
import pandas as pd # manipulate tabular data
import nibabel as nib # load the beta maps in python
from nilearn import plotting # plot nifti images
Download relevant data from ds000164 (and Atlas Files)¶
The subject data came from openneuro [notebook-1]. The atlas data came from a recently published parcellation in a publically accessible github repository.
# atlas github repo for reference:
"""https://github.com/ThomasYeoLab/CBIG/raw/master/stable_projects/\
brain_parcellation/Schaefer2018_LocalGlobal/Parcellations/MNI/"""
data_dir = tempfile.mkdtemp()
print('Our working directory: {}'.format(data_dir))
# download the tar data
url = "https://www.dropbox.com/s/fvtyld08srwl3x9/ds000164-test_v2.tar.gz?dl=1"
tar_file = os.path.join(data_dir, "ds000164.tar.gz")
u = urllib.request.urlopen(url)
data = u.read()
u.close()
# write tar data to file
with open(tar_file, "wb") as f:
f.write(data)
# extract the data
tar = tarfile.open(tar_file, mode='r|gz')
tar.extractall(path=data_dir)
os.remove(tar_file)
Out:
Our working directory: /tmp/tmp2_si98o3
Display the minimal dataset necessary to run nibs¶
# https://stackoverflow.com/questions/9727673/list-directory-tree-structure-in-python
def list_files(startpath):
for root, dirs, files in os.walk(startpath):
level = root.replace(startpath, '').count(os.sep)
indent = ' ' * 4 * (level)
print('{}{}/'.format(indent, os.path.basename(root)))
subindent = ' ' * 4 * (level + 1)
for f in files:
print('{}{}'.format(subindent, f))
list_files(data_dir)
Out:
tmp2_si98o3/
ds000164/
T1w.json
README
task-stroop_bold.json
dataset_description.json
task-stroop_events.json
CHANGES
derivatives/
data/
Schaefer2018_100Parcels_7Networks_order.txt
Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz
fmriprep/
dataset_description.json
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
sub-001_task-stroop_desc-confounds_regressors.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-brain_mask.nii.gz
sub-001/
anat/
sub-001_T1w.nii.gz
func/
sub-001_task-stroop_bold.nii.gz
sub-001_task-stroop_events.tsv
Manipulate events file so it satifies assumptions¶
1. the correct column has 1’s and 0’s corresponding to correct and incorrect, respectively. 2. the condition column is renamed to trial_type nibs currently depends on the “correct” column being binary and the “trial_type” column to contain the trial types of interest.
read the file¶
events_file = os.path.join(data_dir,
"ds000164",
"sub-001",
"func",
"sub-001_task-stroop_events.tsv")
events_df = pd.read_csv(events_file, sep='\t', na_values="n/a")
print(events_df.head())
Out:
onset duration correct condition response_time
0 0.342 1 Y neutral 1.186
1 3.345 1 Y congruent 0.667
2 12.346 1 Y congruent 0.614
3 15.349 1 Y neutral 0.696
4 18.350 1 Y neutral 0.752
replace condition with trial_type¶
events_df.rename({"condition": "trial_type"}, axis='columns', inplace=True)
print(events_df.head())
Out:
onset duration correct trial_type response_time
0 0.342 1 Y neutral 1.186
1 3.345 1 Y congruent 0.667
2 12.346 1 Y congruent 0.614
3 15.349 1 Y neutral 0.696
4 18.350 1 Y neutral 0.752
save the file¶
events_df.to_csv(events_file, sep="\t", na_rep="n/a", index=False)
Manipulate the region order file¶
There are several adjustments to the atlas file that need to be completed before we can pass it into nibs. Importantly, the relevant column names MUST be named “index” and “regions”. “index” refers to which integer within the file corresponds to which region in the atlas nifti file. “regions” refers the name of each region in the atlas nifti file.
read the atlas file¶
atlas_txt = os.path.join(data_dir,
"ds000164",
"derivatives",
"data",
"Schaefer2018_100Parcels_7Networks_order.txt")
atlas_df = pd.read_csv(atlas_txt, sep="\t", header=None)
print(atlas_df.head())
Out:
0 1 2 3 4 5
0 1 7Networks_LH_Vis_1 120 18 131 0
1 2 7Networks_LH_Vis_2 120 18 132 0
2 3 7Networks_LH_Vis_3 120 18 133 0
3 4 7Networks_LH_Vis_4 120 18 135 0
4 5 7Networks_LH_Vis_5 120 18 136 0
drop color coding columns¶
atlas_df.drop([2, 3, 4, 5], axis='columns', inplace=True)
print(atlas_df.head())
Out:
0 1
0 1 7Networks_LH_Vis_1
1 2 7Networks_LH_Vis_2
2 3 7Networks_LH_Vis_3
3 4 7Networks_LH_Vis_4
4 5 7Networks_LH_Vis_5
rename columns with the approved headings: “index” and “regions”¶
atlas_df.rename({0: 'index', 1: 'regions'}, axis='columns', inplace=True)
print(atlas_df.head())
Out:
index regions
0 1 7Networks_LH_Vis_1
1 2 7Networks_LH_Vis_2
2 3 7Networks_LH_Vis_3
3 4 7Networks_LH_Vis_4
4 5 7Networks_LH_Vis_5
remove prefix “7Networks”¶
atlas_df.replace(regex={'7Networks_(.*)': '\\1'}, inplace=True)
print(atlas_df.head())
Out:
index regions
0 1 LH_Vis_1
1 2 LH_Vis_2
2 3 LH_Vis_3
3 4 LH_Vis_4
4 5 LH_Vis_5
write out the file as .tsv¶
atlas_tsv = atlas_txt.replace(".txt", ".tsv")
atlas_df.to_csv(atlas_tsv, sep="\t", index=False)
Run nibs¶
This demonstration mimics how you would use nibs on the command line If you only wanted the beta maps and not the correlation matrices, do not include the atlas (-a) and lookup table options (-l)
out_dir = os.path.join(data_dir, "ds000164", "derivatives")
work_dir = os.path.join(out_dir, "work")
atlas_mni_file = os.path.join(data_dir,
"ds000164",
"derivatives",
"data",
"Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz")
cmd = """\
nibs -c WhiteMatter CSF \
--participant-label 001 \
--estimator lsa \
--hrf-model glover \
-w {work_dir} \
-a {atlas_mni_file} \
-l {atlas_tsv} \
{bids_dir} \
fmriprep \
{out_dir} \
participant
""".format(atlas_mni_file=atlas_mni_file,
atlas_tsv=atlas_tsv,
bids_dir=os.path.join(data_dir, "ds000164"),
out_dir=out_dir,
work_dir=work_dir)
# Since we cannot run bash commands inside this tutorial
# we are printing the actual bash command so you can see it
# in the output
print("The Example Command:\n", cmd)
# call nibs
p = Popen(cmd, shell=True, stdout=PIPE, stderr=STDOUT)
while True:
line = p.stdout.readline()
if not line:
break
print(line)
Out:
The Example Command:
nibs -c WhiteMatter CSF --participant-label 001 --estimator lsa --hrf-model glover -w /tmp/tmp2_si98o3/ds000164/derivatives/work -a /tmp/tmp2_si98o3/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz -l /tmp/tmp2_si98o3/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order.tsv /tmp/tmp2_si98o3/ds000164 fmriprep /tmp/tmp2_si98o3/ds000164/derivatives participant
b'200124-05:13:38,364 nipype.utils INFO:\n'
b'\t Running nipype version 1.3.2 (latest: 1.4.0)\n'
b'200124-05:13:38,412 nipype.workflow INFO:\n'
b"\t Workflow nibetaseries_participant_wf settings: ['check', 'execution', 'logging', 'monitoring']\n"
b'200124-05:13:38,422 nipype.workflow INFO:\n'
b'\t Running in parallel.\n'
b'200124-05:13:38,426 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/bids/layout/models.py:157: UserWarning: Accessing entities as attributes is deprecated as of 0.7. Please use the .entities dictionary instead (i.e., .entities['task'] instead of .task.\n"
b' % (attr, attr))\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/bids/layout/models.py:157: UserWarning: Accessing entities as attributes is deprecated as of 0.7. Please use the .entities dictionary instead (i.e., .entities['space'] instead of .space.\n"
b' % (attr, attr))\n'
b'200124-05:13:38,477 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/3901a92fcfd79d40c4b452de2ef824046b034750/betaseries_node".\n'
b'200124-05:13:38,483 nipype.workflow INFO:\n'
b'\t [Node] Running "betaseries_node" ("nibetaseries.interfaces.nistats.LSABetaSeries")\n'
b'200124-05:13:40,427 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 1 tasks, and 0 jobs ready. Free memory (GB): 6.81/7.01, Free processors: 3/4.\n'
b' Currently running:\n'
b' * nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node\n'
b'200124-05:13:44,978 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node".\n'
b'200124-05:13:46,431 nipype.workflow INFO:\n'
b'\t [Job 0] Completed (nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node).\n'
b'200124-05:13:46,433 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 2 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:13:46,474 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.check_beta_series_list" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/check_beta_series_list".\n'
b'200124-05:13:46,480 nipype.workflow INFO:\n'
b'\t [Node] Running "check_beta_series_list" ("nipype.interfaces.utility.wrappers.Function")\n'
b'200124-05:13:46,491 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.check_beta_series_list".\n'
b'200124-05:13:48,433 nipype.workflow INFO:\n'
b'\t [Job 2] Completed (nibetaseries_participant_wf.single_subject001_wf.check_beta_series_list).\n'
b'200124-05:13:48,435 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 4 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:13:48,481 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file0".\n'
b'200124-05:13:48,482 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file1".\n'
b'200124-05:13:48,485 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:48,485 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:48,488 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file2".\n'
b'200124-05:13:48,490 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:48,500 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file2".\n'
b'200124-05:13:48,500 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file0".\n'
b'200124-05:13:48,500 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file1".\n'
b'200124-05:13:50,435 nipype.workflow INFO:\n'
b'\t [Job 6] Completed (_ds_betaseries_file0).\n'
b'200124-05:13:50,436 nipype.workflow INFO:\n'
b'\t [Job 7] Completed (_ds_betaseries_file1).\n'
b'200124-05:13:50,436 nipype.workflow INFO:\n'
b'\t [Job 8] Completed (_ds_betaseries_file2).\n'
b'200124-05:13:50,438 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 4 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:13:50,480 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file".\n'
b'200124-05:13:50,482 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node1".\n'
b'200124-05:13:50,483 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node2".\n'
b'200124-05:13:50,484 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node1" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'200124-05:13:50,485 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file0".\n'
b'200124-05:13:50,485 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node2" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'200124-05:13:50,486 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node0".\n'
b'200124-05:13:50,488 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:50,489 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node0" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'200124-05:13:50,501 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file0".\n'
b'200124-05:13:50,502 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file1".\n'
b'200124-05:13:50,505 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:50,514 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file1".\n'
b'200124-05:13:50,515 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_betaseries_file/mapflow/_ds_betaseries_file2".\n'
b'200124-05:13:50,518 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:13:50,527 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file2".\n'
b'200124-05:13:50,529 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file".\n'
b'200124-05:13:52,437 nipype.workflow INFO:\n'
b'\t [Job 1] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file).\n'
b'200124-05:13:52,438 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 3 tasks, and 0 jobs ready. Free memory (GB): 6.41/7.01, Free processors: 1/4.\n'
b' Currently running:\n'
b' * _atlas_corr_node2\n'
b' * _atlas_corr_node1\n'
b' * _atlas_corr_node0\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp2_si98o3/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/3901a92fcfd79d40c4b452de2ef824046b034750/betaseries_node/desc-neutral_betase\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'200124-05:13:58,803 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node0".\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp2_si98o3/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/3901a92fcfd79d40c4b452de2ef824046b034750/betaseries_node/desc-incongruent_be\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'200124-05:13:59,308 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node2".\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp2_si98o3/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/3901a92fcfd79d40c4b452de2ef824046b034750/betaseries_node/desc-congruent_beta\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'200124-05:13:59,771 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node1".\n'
b'200124-05:14:00,445 nipype.workflow INFO:\n'
b'\t [Job 9] Completed (_atlas_corr_node0).\n'
b'200124-05:14:00,445 nipype.workflow INFO:\n'
b'\t [Job 10] Completed (_atlas_corr_node1).\n'
b'200124-05:14:00,446 nipype.workflow INFO:\n'
b'\t [Job 11] Completed (_atlas_corr_node2).\n'
b'200124-05:14:00,447 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:14:00,480 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node".\n'
b'200124-05:14:00,483 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node0".\n'
b'200124-05:14:00,485 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node0" - collecting precomputed outputs\n'
b'200124-05:14:00,485 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node0" found cached.\n'
b'200124-05:14:00,486 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node1".\n'
b'200124-05:14:00,488 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node1" - collecting precomputed outputs\n'
b'200124-05:14:00,488 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node1" found cached.\n'
b'200124-05:14:00,489 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/3901a92fcfd79d40c4b452de2ef824046b034750/atlas_corr_node/mapflow/_atlas_corr_node2".\n'
b'200124-05:14:00,490 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node2" - collecting precomputed outputs\n'
b'200124-05:14:00,490 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node2" found cached.\n'
b'200124-05:14:00,491 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node".\n'
b'200124-05:14:02,447 nipype.workflow INFO:\n'
b'\t [Job 3] Completed (nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node).\n'
b'200124-05:14:02,449 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 2 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:14:04,450 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 6 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:14:04,479 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig0".\n'
b'200124-05:14:04,480 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig1".\n'
b'200124-05:14:04,481 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:04,481 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig2".\n'
b'200124-05:14:04,482 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:04,483 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:04,488 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix0".\n'
b'200124-05:14:04,490 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:04,492 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig0".\n'
b'200124-05:14:04,492 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig1".\n'
b'200124-05:14:04,493 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig2".\n'
b'200124-05:14:04,499 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix0".\n'
b'200124-05:14:06,452 nipype.workflow INFO:\n'
b'\t [Job 12] Completed (_ds_correlation_fig0).\n'
b'200124-05:14:06,452 nipype.workflow INFO:\n'
b'\t [Job 13] Completed (_ds_correlation_fig1).\n'
b'200124-05:14:06,453 nipype.workflow INFO:\n'
b'\t [Job 14] Completed (_ds_correlation_fig2).\n'
b'200124-05:14:06,453 nipype.workflow INFO:\n'
b'\t [Job 15] Completed (_ds_correlation_matrix0).\n'
b'200124-05:14:06,455 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 3 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:14:06,491 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig".\n'
b'200124-05:14:06,491 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix1".\n'
b'200124-05:14:06,492 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix2".\n'
b'200124-05:14:06,493 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:06,494 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:06,495 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig0".\n'
b'200124-05:14:06,498 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:06,501 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix1".\n'
b'200124-05:14:06,502 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix2".\n'
b'200124-05:14:06,507 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig0".\n'
b'200124-05:14:06,508 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig1".\n'
b'200124-05:14:06,510 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:06,520 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig1".\n'
b'200124-05:14:06,521 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_fig/mapflow/_ds_correlation_fig2".\n'
b'200124-05:14:06,523 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:06,534 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig2".\n'
b'200124-05:14:06,535 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig".\n'
b'200124-05:14:08,454 nipype.workflow INFO:\n'
b'\t [Job 4] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig).\n'
b'200124-05:14:08,455 nipype.workflow INFO:\n'
b'\t [Job 16] Completed (_ds_correlation_matrix1).\n'
b'200124-05:14:08,455 nipype.workflow INFO:\n'
b'\t [Job 17] Completed (_ds_correlation_matrix2).\n'
b'200124-05:14:08,456 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'200124-05:14:08,491 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix".\n'
b'200124-05:14:08,494 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix0" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix0".\n'
b'200124-05:14:08,496 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:08,503 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix0".\n'
b'200124-05:14:08,504 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix1" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix1".\n'
b'200124-05:14:08,506 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:08,513 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix1".\n'
b'200124-05:14:08,514 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix2" in "/tmp/tmp2_si98o3/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/3901a92fcfd79d40c4b452de2ef824046b034750/ds_correlation_matrix/mapflow/_ds_correlation_matrix2".\n'
b'200124-05:14:08,515 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'200124-05:14:08,521 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix2".\n'
b'200124-05:14:08,522 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix".\n'
b'200124-05:14:10,456 nipype.workflow INFO:\n'
b'\t [Job 5] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix).\n'
b'200124-05:14:10,458 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 0 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/nibetaseries/interfaces/nistats.py:180: DeprecationWarning: The parameter "mask" will be removed in next release of Nistats. Please use the parameter "mask_img" instead.\n'
b' verbose=1\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 4 seconds\n'
b'\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b'pandoc: Error running filter pandoc-citeproc:\n'
b"Could not find executable 'pandoc-citeproc'.\n"
b'Could not generate CITATION.html file:\n'
b'pandoc -s --bibliography /home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/nibetaseries/data/references.bib --filter pandoc-citeproc --metadata pagetitle="NiBetaSeries citation boilerplate" /tmp/tmp2_si98o3/ds000164/derivatives/nibetaseries/logs/CITATION.md -o /tmp/tmp2_si98o3/ds000164/derivatives/nibetaseries/logs/CITATION.html\n'
Observe generated outputs¶
list_files(data_dir)
Out:
tmp2_si98o3/
ds000164/
T1w.json
README
task-stroop_bold.json
dataset_description.json
task-stroop_events.json
CHANGES
derivatives/
nibetaseries/
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_correlation.svg
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_correlation.svg
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_correlation.svg
logs/
CITATION.tex
CITATION.md
CITATION.bib
data/
Schaefer2018_100Parcels_7Networks_order.tsv
Schaefer2018_100Parcels_7Networks_order.txt
Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz
work/
dbcache
NiBetaSeries_work/
nibetaseries_participant_wf/
graph1.json
d3.js
index.html
graph.json
single_subject001_wf/
correlation_wf/
3901a92fcfd79d40c4b452de2ef824046b034750/
atlas_corr_node/
_node.pklz
result_atlas_corr_node.pklz
_inputs.pklz
_0xcb1f362dddb54d2fc35e5825ba16448b.json
_report/
report.rst
mapflow/
_atlas_corr_node2/
_node.pklz
_inputs.pklz
desc-incongruent_correlation.tsv
desc-incongruent_correlation.svg
_0x5e1f7323b96ea30157e49866846d090d.json
result__atlas_corr_node2.pklz
_report/
report.rst
_atlas_corr_node0/
_node.pklz
_inputs.pklz
desc-neutral_correlation.svg
result__atlas_corr_node0.pklz
_0x5a60ff19c6a19d9bad170136fa9ee816.json
desc-neutral_correlation.tsv
_report/
report.rst
_atlas_corr_node1/
_node.pklz
_inputs.pklz
desc-congruent_correlation.svg
desc-congruent_correlation.tsv
_0x16c0fc73fec275e3d8021b284caba70b.json
result__atlas_corr_node1.pklz
_report/
report.rst
3901a92fcfd79d40c4b452de2ef824046b034750/
check_beta_series_list/
_node.pklz
_0x3ddcf8e1f8869833d3373ca792dffca5.json
_inputs.pklz
result_check_beta_series_list.pklz
_report/
report.rst
ds_correlation_matrix/
_node.pklz
result_ds_correlation_matrix.pklz
_inputs.pklz
_0x05aeb5151f9ebe01b9a9137258470b2a.json
_report/
report.rst
mapflow/
_ds_correlation_matrix0/
_0xd1cf1953f001a382796360b0f23d7db8.json
_node.pklz
result__ds_correlation_matrix0.pklz
_inputs.pklz
_report/
report.rst
_ds_correlation_matrix1/
_node.pklz
_inputs.pklz
_0x5f7fa35ab4342e872965c1cb8cfe6e98.json
result__ds_correlation_matrix1.pklz
_report/
report.rst
_ds_correlation_matrix2/
_node.pklz
_0x803d436706ee4d348c1ab725f6875c00.json
_inputs.pklz
result__ds_correlation_matrix2.pklz
_report/
report.rst
ds_correlation_fig/
_node.pklz
_inputs.pklz
result_ds_correlation_fig.pklz
_0xe230b37a838ad69c74ca5fd6ee29309f.json
_report/
report.rst
mapflow/
_ds_correlation_fig0/
_node.pklz
_inputs.pklz
result__ds_correlation_fig0.pklz
_0x597f5c4a21553cee15000f440dda773d.json
_report/
report.rst
_ds_correlation_fig2/
_0x7976abdcfe59071f02f3cf1102a91656.json
_node.pklz
_inputs.pklz
result__ds_correlation_fig2.pklz
_report/
report.rst
_ds_correlation_fig1/
_node.pklz
_inputs.pklz
result__ds_correlation_fig1.pklz
_0x2f98a5303b005be0df4fb687694c7731.json
_report/
report.rst
ds_betaseries_file/
_node.pklz
_inputs.pklz
_0x5ff15d098d6736c64da912d86c78dc17.json
result_ds_betaseries_file.pklz
_report/
report.rst
mapflow/
_ds_betaseries_file1/
_node.pklz
_inputs.pklz
result__ds_betaseries_file1.pklz
_0x3d84861b40f3cec9c3fe80a69be65e23.json
_report/
report.rst
_ds_betaseries_file0/
_node.pklz
_inputs.pklz
result__ds_betaseries_file0.pklz
_0x465cff8852beb4621de39e3b345ba1c8.json
_report/
report.rst
_ds_betaseries_file2/
_node.pklz
_inputs.pklz
result__ds_betaseries_file2.pklz
_0x293b434daae39dfa5f78b81456e7ff96.json
_report/
report.rst
betaseries_wf/
3901a92fcfd79d40c4b452de2ef824046b034750/
betaseries_node/
desc-neutral_betaseries.nii.gz
_node.pklz
_inputs.pklz
_0xc338cae97b2926f8302b1fa116bce156.json
result_betaseries_node.pklz
desc-congruent_betaseries.nii.gz
desc-incongruent_betaseries.nii.gz
_report/
report.rst
fmriprep/
fMRIPrep.sqlite
dataset_description.json
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
sub-001_task-stroop_desc-confounds_regressors.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-brain_mask.nii.gz
sub-001/
anat/
sub-001_T1w.nii.gz
func/
sub-001_task-stroop_bold.nii.gz
sub-001_task-stroop_events.tsv
Collect correlation results¶
output_path = os.path.join(out_dir, "nibetaseries", "sub-001", "func")
trial_types = ['congruent', 'incongruent', 'neutral']
filename_template = ('sub-001_task-stroop_space-MNI152NLin2009cAsym_'
'desc-{trial_type}_{suffix}.{ext}')
pd_dict = {}
for trial_type in trial_types:
fname = filename_template.format(trial_type=trial_type, suffix='correlation', ext='tsv')
file_path = os.path.join(output_path, fname)
pd_dict[trial_type] = pd.read_csv(file_path, sep='\t', na_values="n/a", index_col=0)
# display example matrix
print(pd_dict[trial_type].head())
Out:
LH_Vis_1 LH_Vis_2 ... RH_Default_PCC_1 RH_Default_PCC_2
LH_Vis_1 NaN 0.218441 ... -0.229994 0.030440
LH_Vis_2 0.218441 NaN ... -0.272071 0.051983
LH_Vis_3 -0.193264 0.448418 ... 0.087879 0.083368
LH_Vis_4 0.122960 0.526585 ... -0.185481 0.188477
LH_Vis_5 0.284347 0.534536 ... -0.393536 -0.288098
[5 rows x 100 columns]
Graph the correlation results¶
fig, axes = plt.subplots(nrows=3, ncols=1, sharex=True, sharey=True, figsize=(10, 30),
gridspec_kw={'wspace': 0.025, 'hspace': 0.075})
cbar_ax = fig.add_axes([.91, .3, .03, .4])
r = 0
for trial_type, df in pd_dict.items():
g = sns.heatmap(df, ax=axes[r], vmin=-.5, vmax=1., square=True,
cbar=True, cbar_ax=cbar_ax)
axes[r].set_title(trial_type)
# iterate over rows
r += 1
plt.tight_layout()
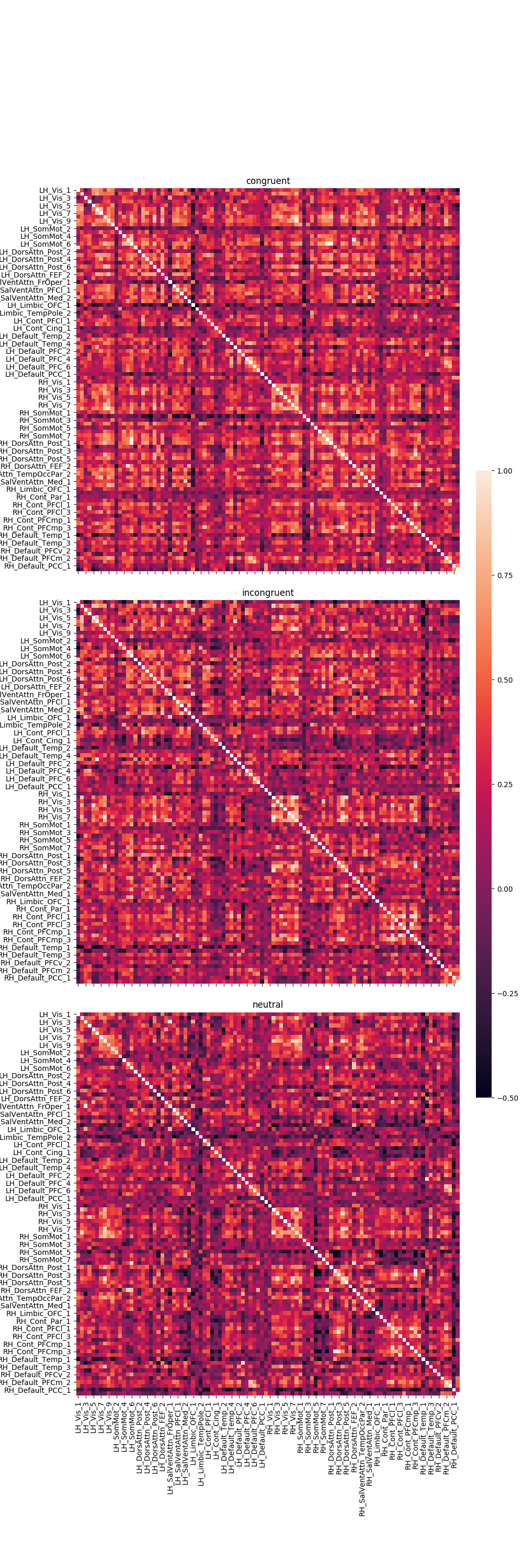
Out:
/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.2/lib/python3.7/site-packages/matplotlib/figure.py:2299: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
warnings.warn("This figure includes Axes that are not compatible "
Collect beta map results¶
nii_dict = {}
for trial_type in trial_types:
fname = filename_template.format(trial_type=trial_type, suffix='betaseries', ext='nii.gz')
file_path = os.path.join(output_path, fname)
nii_dict[trial_type] = nib.load(file_path)
# view incongruent beta_maps
nib.viewers.OrthoSlicer3D(nii_dict['incongruent'].get_fdata(),
title="Incongruent Betas").set_position(10, 13, 10)
Graph beta map standard deviation¶
We can find where the betas have the highest standard deviation for each trial type. Unsuprisingly, the largest deviations are near the edge of the brain mask and the subcortical regions.
# standard deviations for each trial type
std_dict = {tt: nib.Nifti1Image(img.get_fdata().std(axis=-1), img.affine, img.header)
for tt, img in nii_dict.items()}
fig, axes = plt.subplots(nrows=3, ncols=1, figsize=(10, 20))
for idx, (trial_type, nii) in enumerate(std_dict.items()):
plotting.plot_stat_map(nii, title=trial_type, cut_coords=(0, 0, 0),
threshold=5, vmax=20, axes=axes[idx])
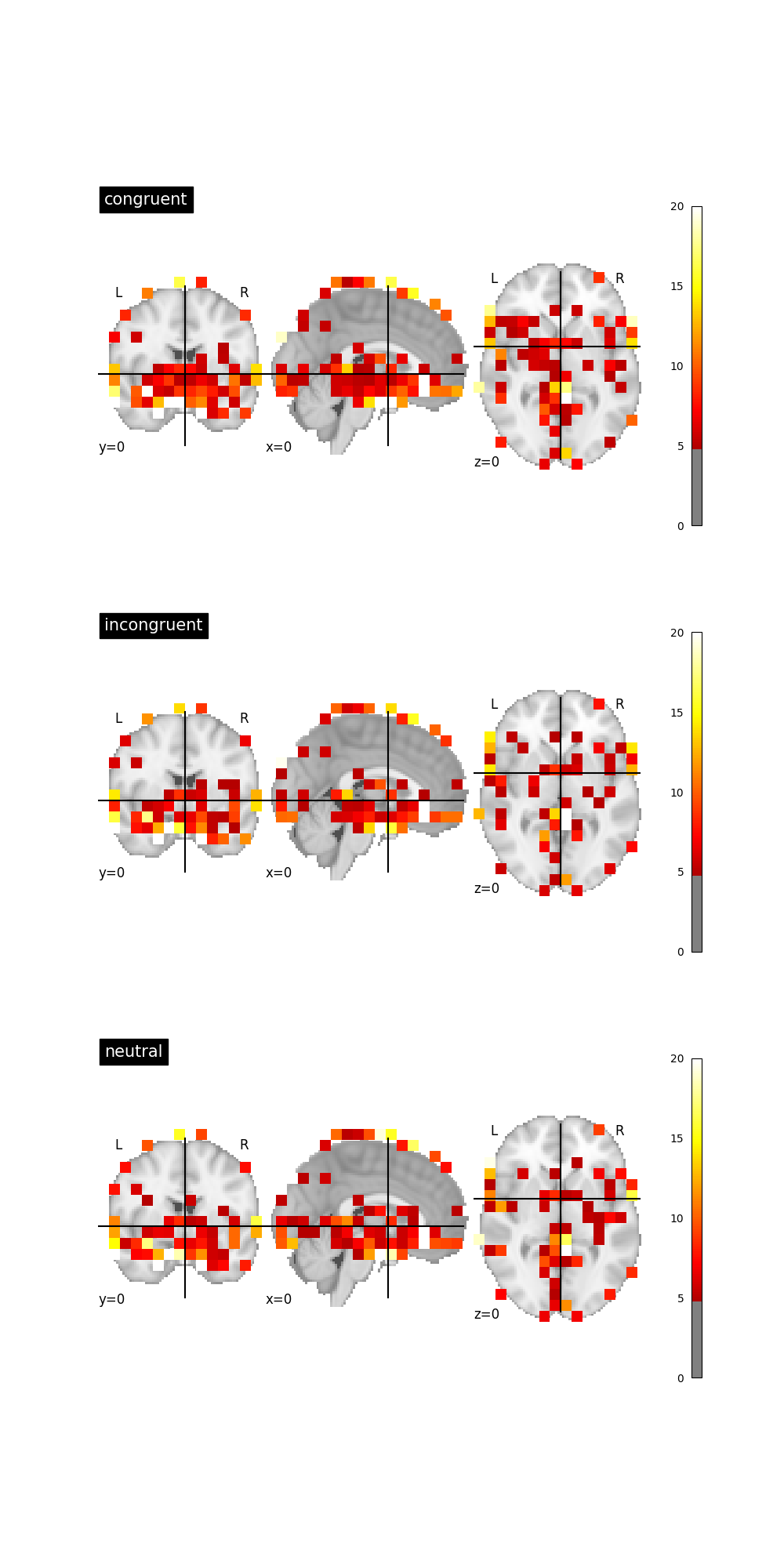
References¶
- notebook-1
Timothy D Verstynen. The organization and dynamics of corticostriatal pathways link the medial orbitofrontal cortex to future behavioral responses. Journal of Neurophysiology, 112(10):2457–2469, 2014. URL: https://doi.org/10.1152/jn.00221.2014, doi:10.1152/jn.00221.2014.
Total running time of the script: ( 0 minutes 49.341 seconds)