Note
Click here to download the full example code or run this example in your browser via Binder
Running NiBetaSeries¶
This example runs through a basic call of NiBetaSeries using
the commandline entry point nibs.
While this example is using python, typically nibs will be
called directly on the commandline.
Import all the necessary packages¶
import tempfile # make a temporary directory for files
import os # interact with the filesystem
import urllib.request # grad data from internet
import tarfile # extract files from tar
from subprocess import Popen, PIPE, STDOUT # enable calling commandline
import matplotlib.pyplot as plt # manipulate figures
import seaborn as sns # display results
import pandas as pd # manipulate tabular data
Download relevant data from ds000164 (and Atlas Files)¶
The subject data came from openneuro [notebook-1]. The atlas data came from a recently published parcellation in a publically accessible github repository.
# atlas github repo for reference:
"""https://github.com/ThomasYeoLab/CBIG/raw/master/stable_projects/\
brain_parcellation/Schaefer2018_LocalGlobal/Parcellations/MNI/"""
data_dir = tempfile.mkdtemp()
print('Our working directory: {}'.format(data_dir))
# download the tar data
url = "https://www.dropbox.com/s/fvtyld08srwl3x9/ds000164-test_v2.tar.gz?dl=1"
tar_file = os.path.join(data_dir, "ds000164.tar.gz")
u = urllib.request.urlopen(url)
data = u.read()
u.close()
# write tar data to file
with open(tar_file, "wb") as f:
f.write(data)
# extract the data
tar = tarfile.open(tar_file, mode='r|gz')
tar.extractall(path=data_dir)
os.remove(tar_file)
Out:
Our working directory: /tmp/tmp4iah86cu
Display the minimal dataset necessary to run nibs¶
# https://stackoverflow.com/questions/9727673/list-directory-tree-structure-in-python
def list_files(startpath):
for root, dirs, files in os.walk(startpath):
level = root.replace(startpath, '').count(os.sep)
indent = ' ' * 4 * (level)
print('{}{}/'.format(indent, os.path.basename(root)))
subindent = ' ' * 4 * (level + 1)
for f in files:
print('{}{}'.format(subindent, f))
list_files(data_dir)
Out:
tmp4iah86cu/
ds000164/
T1w.json
README
task-stroop_bold.json
dataset_description.json
task-stroop_events.json
CHANGES
derivatives/
data/
Schaefer2018_100Parcels_7Networks_order.txt
Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz
fmriprep/
dataset_description.json
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
sub-001_task-stroop_desc-confounds_regressors.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-brain_mask.nii.gz
sub-001/
anat/
sub-001_T1w.nii.gz
func/
sub-001_task-stroop_bold.nii.gz
sub-001_task-stroop_events.tsv
Manipulate events file so it satifies assumptions¶
1. the correct column has 1’s and 0’s corresponding to correct and incorrect, respectively. 2. the condition column is renamed to trial_type nibs currently depends on the “correct” column being binary and the “trial_type” column to contain the trial types of interest.
read the file¶
events_file = os.path.join(data_dir,
"ds000164",
"sub-001",
"func",
"sub-001_task-stroop_events.tsv")
events_df = pd.read_csv(events_file, sep='\t', na_values="n/a")
print(events_df.head())
Out:
onset duration correct condition response_time
0 0.342 1 Y neutral 1.186
1 3.345 1 Y congruent 0.667
2 12.346 1 Y congruent 0.614
3 15.349 1 Y neutral 0.696
4 18.350 1 Y neutral 0.752
replace condition with trial_type¶
events_df.rename({"condition": "trial_type"}, axis='columns', inplace=True)
print(events_df.head())
Out:
onset duration correct trial_type response_time
0 0.342 1 Y neutral 1.186
1 3.345 1 Y congruent 0.667
2 12.346 1 Y congruent 0.614
3 15.349 1 Y neutral 0.696
4 18.350 1 Y neutral 0.752
save the file¶
events_df.to_csv(events_file, sep="\t", na_rep="n/a", index=False)
Manipulate the region order file¶
There are several adjustments to the atlas file that need to be completed before we can pass it into nibs. Importantly, the relevant column names MUST be named “index” and “regions”. “index” refers to which integer within the file corresponds to which region in the atlas nifti file. “regions” refers the name of each region in the atlas nifti file.
read the atlas file¶
atlas_txt = os.path.join(data_dir,
"ds000164",
"derivatives",
"data",
"Schaefer2018_100Parcels_7Networks_order.txt")
atlas_df = pd.read_csv(atlas_txt, sep="\t", header=None)
print(atlas_df.head())
Out:
0 1 2 3 4 5
0 1 7Networks_LH_Vis_1 120 18 131 0
1 2 7Networks_LH_Vis_2 120 18 132 0
2 3 7Networks_LH_Vis_3 120 18 133 0
3 4 7Networks_LH_Vis_4 120 18 135 0
4 5 7Networks_LH_Vis_5 120 18 136 0
drop color coding columns¶
atlas_df.drop([2, 3, 4, 5], axis='columns', inplace=True)
print(atlas_df.head())
Out:
0 1
0 1 7Networks_LH_Vis_1
1 2 7Networks_LH_Vis_2
2 3 7Networks_LH_Vis_3
3 4 7Networks_LH_Vis_4
4 5 7Networks_LH_Vis_5
rename columns with the approved headings: “index” and “regions”¶
atlas_df.rename({0: 'index', 1: 'regions'}, axis='columns', inplace=True)
print(atlas_df.head())
Out:
index regions
0 1 7Networks_LH_Vis_1
1 2 7Networks_LH_Vis_2
2 3 7Networks_LH_Vis_3
3 4 7Networks_LH_Vis_4
4 5 7Networks_LH_Vis_5
remove prefix “7Networks”¶
atlas_df.replace(regex={'7Networks_(.*)': '\\1'}, inplace=True)
print(atlas_df.head())
Out:
index regions
0 1 LH_Vis_1
1 2 LH_Vis_2
2 3 LH_Vis_3
3 4 LH_Vis_4
4 5 LH_Vis_5
write out the file as .tsv¶
atlas_tsv = atlas_txt.replace(".txt", ".tsv")
atlas_df.to_csv(atlas_tsv, sep="\t", index=False)
Run nibs¶
out_dir = os.path.join(data_dir, "ds000164", "derivatives")
work_dir = os.path.join(out_dir, "work")
atlas_mni_file = os.path.join(data_dir,
"ds000164",
"derivatives",
"data",
"Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz")
cmd = """\
nibs -c WhiteMatter CSF \
--participant-label 001 \
--estimator lss \
-w {work_dir} \
-a {atlas_mni_file} \
-l {atlas_tsv} \
{bids_dir} \
fmriprep \
{out_dir} \
participant
""".format(atlas_mni_file=atlas_mni_file,
atlas_tsv=atlas_tsv,
bids_dir=os.path.join(data_dir, "ds000164"),
out_dir=out_dir,
work_dir=work_dir)
# Since we cannot run bash commands inside this tutorial
# we are printing the actual bash command so you can see it
# in the output
print("The Example Command:\n", cmd)
# call nibs
p = Popen(cmd, shell=True, stdout=PIPE, stderr=STDOUT)
while True:
line = p.stdout.readline()
if not line:
break
print(line)
Out:
The Example Command:
nibs -c WhiteMatter CSF --participant-label 001 --estimator lss -w /tmp/tmp4iah86cu/ds000164/derivatives/work -a /tmp/tmp4iah86cu/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz -l /tmp/tmp4iah86cu/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order.tsv /tmp/tmp4iah86cu/ds000164 fmriprep /tmp/tmp4iah86cu/ds000164/derivatives participant
b'191009-00:49:41,502 nipype.workflow INFO:\n'
b"\t Workflow nibetaseries_participant_wf settings: ['check', 'execution', 'logging', 'monitoring']\n"
b'191009-00:49:41,513 nipype.workflow INFO:\n'
b'\t Running in parallel.\n'
b'191009-00:49:41,517 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/bids/layout/models.py:146: UserWarning: Accessing entities as attributes is deprecated as of 0.7. Please use the .entities dictionary instead (i.e., .entities['task'] instead of .task.\n"
b' % (attr, attr))\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/bids/layout/models.py:146: UserWarning: Accessing entities as attributes is deprecated as of 0.7. Please use the .entities dictionary instead (i.e., .entities['space'] instead of .space.\n"
b' % (attr, attr))\n'
b'191009-00:49:41,577 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/betaseries_node".\n'
b'191009-00:49:41,582 nipype.workflow INFO:\n'
b'\t [Node] Running "betaseries_node" ("nibetaseries.interfaces.nistats.LSSBetaSeries")\n'
b'191009-00:49:43,520 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 1 tasks, and 0 jobs ready. Free memory (GB): 6.81/7.01, Free processors: 3/4.\n'
b' Currently running:\n'
b' * nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b"<string>:6: DeprecationWarning: object of type <class 'numpy.float64'> cannot be safely interpreted as an integer.\n"
b"<string>:6: DeprecationWarning: object of type <class 'float'> cannot be safely interpreted as an integer.\n"
b'\n'
b'Computation of 1 runs done in 1 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'191009-00:51:03,980 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node".\n'
b'191009-00:51:05,601 nipype.workflow INFO:\n'
b'\t [Job 0] Completed (nibetaseries_participant_wf.single_subject001_wf.betaseries_wf.betaseries_node).\n'
b'191009-00:51:05,604 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 2 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:07,604 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 6 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:07,644 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file0".\n'
b'191009-00:51:07,645 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file1".\n'
b'191009-00:51:07,647 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:07,648 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:07,655 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file2".\n'
b'191009-00:51:07,658 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:07,662 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node0".\n'
b'191009-00:51:07,664 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node0" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'191009-00:51:07,667 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file0".\n'
b'191009-00:51:07,669 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file1".\n'
b'191009-00:51:07,677 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file2".\n'
b'191009-00:51:09,605 nipype.workflow INFO:\n'
b'\t [Job 5] Completed (_ds_betaseries_file0).\n'
b'191009-00:51:09,606 nipype.workflow INFO:\n'
b'\t [Job 6] Completed (_ds_betaseries_file1).\n'
b'191009-00:51:09,606 nipype.workflow INFO:\n'
b'\t [Job 7] Completed (_ds_betaseries_file2).\n'
b'191009-00:51:09,608 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 1 tasks, and 3 jobs ready. Free memory (GB): 6.81/7.01, Free processors: 3/4.\n'
b' Currently running:\n'
b' * _atlas_corr_node0\n'
b'191009-00:51:09,651 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file".\n'
b'191009-00:51:09,654 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file0".\n'
b'191009-00:51:09,657 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:09,658 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node1".\n'
b'191009-00:51:09,660 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node1" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'191009-00:51:09,666 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node2".\n'
b'191009-00:51:09,668 nipype.workflow INFO:\n'
b'\t [Node] Running "_atlas_corr_node2" ("nibetaseries.interfaces.nilearn.AtlasConnectivity")\n'
b'191009-00:51:09,699 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file0".\n'
b'191009-00:51:09,700 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file1".\n'
b'191009-00:51:09,702 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:09,712 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file1".\n'
b'191009-00:51:09,722 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_betaseries_file2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_betaseries_file/mapflow/_ds_betaseries_file2".\n'
b'191009-00:51:09,725 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_betaseries_file2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:09,736 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_betaseries_file2".\n'
b'191009-00:51:09,747 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file".\n'
b'191009-00:51:11,607 nipype.workflow INFO:\n'
b'\t [Job 1] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_betaseries_file).\n'
b'191009-00:51:11,609 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 3 tasks, and 0 jobs ready. Free memory (GB): 6.41/7.01, Free processors: 1/4.\n'
b' Currently running:\n'
b' * _atlas_corr_node2\n'
b' * _atlas_corr_node1\n'
b' * _atlas_corr_node0\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp4iah86cu/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/betaseries_node/desc-neutral_betase\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'191009-00:51:16,475 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node0".\n'
b'191009-00:51:17,613 nipype.workflow INFO:\n'
b'\t [Job 8] Completed (_atlas_corr_node0).\n'
b'191009-00:51:17,614 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 2 tasks, and 0 jobs ready. Free memory (GB): 6.61/7.01, Free processors: 2/4.\n'
b' Currently running:\n'
b' * _atlas_corr_node2\n'
b' * _atlas_corr_node1\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp4iah86cu/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/betaseries_node/desc-congruent_beta\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'191009-00:51:18,593 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node1".\n'
b'[NiftiLabelsMasker.fit_transform] loading data from /tmp/tmp4iah86cu/ds000164/derivatives/data/Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz\n'
b'Resampling labels\n'
b'[NiftiLabelsMasker.transform_single_imgs] Loading data from /tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/betaseries_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/betaseries_node/desc-incongruent_be\n'
b'[NiftiLabelsMasker.transform_single_imgs] Extracting region signals\n'
b'[NiftiLabelsMasker.transform_single_imgs] Cleaning extracted signals\n'
b'191009-00:51:18,666 nipype.workflow INFO:\n'
b'\t [Node] Finished "_atlas_corr_node2".\n'
b'191009-00:51:19,615 nipype.workflow INFO:\n'
b'\t [Job 9] Completed (_atlas_corr_node1).\n'
b'191009-00:51:19,616 nipype.workflow INFO:\n'
b'\t [Job 10] Completed (_atlas_corr_node2).\n'
b'191009-00:51:19,617 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:19,662 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node".\n'
b'191009-00:51:19,666 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node0".\n'
b'191009-00:51:19,667 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node0" - collecting precomputed outputs\n'
b'191009-00:51:19,667 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node0" found cached.\n'
b'191009-00:51:19,668 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node1".\n'
b'191009-00:51:19,669 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node1" - collecting precomputed outputs\n'
b'191009-00:51:19,669 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node1" found cached.\n'
b'191009-00:51:19,670 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_atlas_corr_node2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/correlation_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/atlas_corr_node/mapflow/_atlas_corr_node2".\n'
b'191009-00:51:19,671 nipype.workflow INFO:\n'
b'\t [Node] Cached "_atlas_corr_node2" - collecting precomputed outputs\n'
b'191009-00:51:19,671 nipype.workflow INFO:\n'
b'\t [Node] "_atlas_corr_node2" found cached.\n'
b'191009-00:51:19,673 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node".\n'
b'191009-00:51:21,617 nipype.workflow INFO:\n'
b'\t [Job 2] Completed (nibetaseries_participant_wf.single_subject001_wf.correlation_wf.atlas_corr_node).\n'
b'191009-00:51:21,620 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 2 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:23,620 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 6 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:23,661 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig0".\n'
b'191009-00:51:23,662 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig1".\n'
b'191009-00:51:23,663 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:23,664 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:23,676 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig1".\n'
b'191009-00:51:23,676 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig0".\n'
b'191009-00:51:23,677 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig2".\n'
b'191009-00:51:23,677 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix0".\n'
b'191009-00:51:23,678 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:23,678 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:23,688 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix0".\n'
b'191009-00:51:23,690 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig2".\n'
b'191009-00:51:25,621 nipype.workflow INFO:\n'
b'\t [Job 11] Completed (_ds_correlation_fig0).\n'
b'191009-00:51:25,622 nipype.workflow INFO:\n'
b'\t [Job 12] Completed (_ds_correlation_fig1).\n'
b'191009-00:51:25,623 nipype.workflow INFO:\n'
b'\t [Job 13] Completed (_ds_correlation_fig2).\n'
b'191009-00:51:25,624 nipype.workflow INFO:\n'
b'\t [Job 14] Completed (_ds_correlation_matrix0).\n'
b'191009-00:51:25,626 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 3 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:25,669 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig".\n'
b'191009-00:51:25,670 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix1".\n'
b'191009-00:51:25,671 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix2".\n'
b'191009-00:51:25,672 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig0".\n'
b'191009-00:51:25,672 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:25,672 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:25,675 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:25,681 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix1".\n'
b'191009-00:51:25,682 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix2".\n'
b'191009-00:51:25,687 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig0".\n'
b'191009-00:51:25,688 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig1".\n'
b'191009-00:51:25,690 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:25,702 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig1".\n'
b'191009-00:51:25,703 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_fig2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_fig/mapflow/_ds_correlation_fig2".\n'
b'191009-00:51:25,705 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_fig2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:25,717 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_fig2".\n'
b'191009-00:51:25,719 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig".\n'
b'191009-00:51:27,623 nipype.workflow INFO:\n'
b'\t [Job 3] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_correlation_fig).\n'
b'191009-00:51:27,625 nipype.workflow INFO:\n'
b'\t [Job 15] Completed (_ds_correlation_matrix1).\n'
b'191009-00:51:27,625 nipype.workflow INFO:\n'
b'\t [Job 16] Completed (_ds_correlation_matrix2).\n'
b'191009-00:51:27,627 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'191009-00:51:27,678 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix".\n'
b'191009-00:51:27,681 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix0" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix0".\n'
b'191009-00:51:27,683 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix0" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:27,693 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix0".\n'
b'191009-00:51:27,694 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix1" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix1".\n'
b'191009-00:51:27,696 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix1" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:27,706 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix1".\n'
b'191009-00:51:27,707 nipype.workflow INFO:\n'
b'\t [Node] Setting-up "_ds_correlation_matrix2" in "/tmp/tmp4iah86cu/ds000164/derivatives/work/NiBetaSeries_work/nibetaseries_participant_wf/single_subject001_wf/dfb9084b801be401880a15ffb747f5a5a15543c2/ds_correlation_matrix/mapflow/_ds_correlation_matrix2".\n'
b'191009-00:51:27,709 nipype.workflow INFO:\n'
b'\t [Node] Running "_ds_correlation_matrix2" ("nibetaseries.interfaces.bids.DerivativesDataSink")\n'
b'191009-00:51:27,718 nipype.workflow INFO:\n'
b'\t [Node] Finished "_ds_correlation_matrix2".\n'
b'191009-00:51:27,719 nipype.workflow INFO:\n'
b'\t [Node] Finished "nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix".\n'
b'191009-00:51:29,625 nipype.workflow INFO:\n'
b'\t [Job 4] Completed (nibetaseries_participant_wf.single_subject001_wf.ds_correlation_matrix).\n'
b'191009-00:51:29,627 nipype.workflow INFO:\n'
b'\t [MultiProc] Running 0 tasks, and 0 jobs ready. Free memory (GB): 7.01/7.01, Free processors: 4/4.\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'Computing run 1 out of 1 runs (go take a coffee, a big one)\n'
b'\n'
b'Computation of 1 runs done in 0 seconds\n'
b'\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/matplotlib/cbook/__init__.py:2366: DeprecationWarning: Using or importing the ABCs from 'collections' instead of from 'collections.abc' is deprecated, and in 3.8 it will stop working\n"
b' return list(data) if isinstance(data, collections.MappingView) else data\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/matplotlib/cbook/__init__.py:2366: DeprecationWarning: Using or importing the ABCs from 'collections' instead of from 'collections.abc' is deprecated, and in 3.8 it will stop working\n"
b' return list(data) if isinstance(data, collections.MappingView) else data\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: RuntimeWarning: numpy.ufunc size changed, may indicate binary incompatibility. Expected 216, got 192\n'
b' return f(*args, **kwds)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/importlib/_bootstrap.py:219: ImportWarning: can't resolve package from __spec__ or __package__, falling back on __name__ and __path__\n"
b' return f(*args, **kwds)\n'
b'/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/nibetaseries/interfaces/nilearn.py:85: RuntimeWarning: invalid value encountered in greater\n'
b' n_lines = int(np.sum(connmat > 0) / 2)\n'
b"/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/matplotlib/cbook/__init__.py:2366: DeprecationWarning: Using or importing the ABCs from 'collections' instead of from 'collections.abc' is deprecated, and in 3.8 it will stop working\n"
b' return list(data) if isinstance(data, collections.MappingView) else data\n'
b'pandoc: Error running filter pandoc-citeproc:\n'
b"Could not find executable 'pandoc-citeproc'.\n"
b'Could not generate CITATION.html file:\n'
b'pandoc -s --bibliography /home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/nibetaseries/data/references.bib --filter pandoc-citeproc --metadata pagetitle="NiBetaSeries citation boilerplate" /tmp/tmp4iah86cu/ds000164/derivatives/nibetaseries/logs/CITATION.md -o /tmp/tmp4iah86cu/ds000164/derivatives/nibetaseries/logs/CITATION.html\n'
Observe generated outputs¶
list_files(data_dir)
Out:
tmp4iah86cu/
ds000164/
T1w.json
README
task-stroop_bold.json
dataset_description.json
task-stroop_events.json
CHANGES
derivatives/
nibetaseries/
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_correlation.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-neutral_correlation.svg
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_betaseries.nii.gz
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-congruent_correlation.svg
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-incongruent_correlation.svg
logs/
CITATION.tex
CITATION.md
CITATION.bib
data/
Schaefer2018_100Parcels_7Networks_order.tsv
Schaefer2018_100Parcels_7Networks_order.txt
Schaefer2018_100Parcels_7Networks_order_FSLMNI152_2mm.nii.gz
work/
NiBetaSeries_work/
nibetaseries_participant_wf/
graph1.json
d3.js
index.html
graph.json
single_subject001_wf/
dfb9084b801be401880a15ffb747f5a5a15543c2/
ds_correlation_matrix/
_node.pklz
result_ds_correlation_matrix.pklz
_inputs.pklz
_0x5ffd430f3de3efe501bc13ee709ea7ad.json
_report/
report.rst
mapflow/
_ds_correlation_matrix0/
_node.pklz
result__ds_correlation_matrix0.pklz
_inputs.pklz
_0xae64769425e796a3c9adb18d4fbe0b86.json
_report/
report.rst
_ds_correlation_matrix1/
_node.pklz
_inputs.pklz
_0xdbb1a8dc619aa2a8be441808620b14e6.json
result__ds_correlation_matrix1.pklz
_report/
report.rst
_ds_correlation_matrix2/
_node.pklz
_0x1eb7aa04af561118543d124f4395210b.json
_inputs.pklz
result__ds_correlation_matrix2.pklz
_report/
report.rst
ds_correlation_fig/
_node.pklz
_inputs.pklz
result_ds_correlation_fig.pklz
_0x3525f525f193d5d191a342823e53e97e.json
_report/
report.rst
mapflow/
_ds_correlation_fig0/
_node.pklz
_0x0c3357a2eaba7b19b11fd75c6b354eac.json
_inputs.pklz
result__ds_correlation_fig0.pklz
_report/
report.rst
_ds_correlation_fig2/
_node.pklz
_inputs.pklz
result__ds_correlation_fig2.pklz
_0x5635b708e0d61594527df89ef5bcac48.json
_report/
report.rst
_ds_correlation_fig1/
_node.pklz
_inputs.pklz
result__ds_correlation_fig1.pklz
_0x6273d11ebe0d3c9b85ecdf5314fb4675.json
_report/
report.rst
ds_betaseries_file/
_node.pklz
_inputs.pklz
_0x690495b6052bca8306a9e3f3248da8f4.json
result_ds_betaseries_file.pklz
_report/
report.rst
mapflow/
_ds_betaseries_file1/
_0x592c926441f32453e9c4e64e6af84355.json
_node.pklz
_inputs.pklz
result__ds_betaseries_file1.pklz
_report/
report.rst
_ds_betaseries_file0/
_node.pklz
_inputs.pklz
_0x59e3f0527e629f699304198e181e10d0.json
result__ds_betaseries_file0.pklz
_report/
report.rst
_ds_betaseries_file2/
_0x4a1fe41414057f7b84bc6dc3890ed9ef.json
_node.pklz
_inputs.pklz
result__ds_betaseries_file2.pklz
_report/
report.rst
correlation_wf/
dfb9084b801be401880a15ffb747f5a5a15543c2/
atlas_corr_node/
_node.pklz
result_atlas_corr_node.pklz
_inputs.pklz
_0xcac1e4905b9915f4daf625d44f6442c0.json
_report/
report.rst
mapflow/
_atlas_corr_node2/
_node.pklz
_inputs.pklz
desc-incongruent_correlation.tsv
_0x63c4c138db79321588cb4f27c5273e31.json
desc-incongruent_correlation.svg
result__atlas_corr_node2.pklz
_report/
report.rst
_atlas_corr_node0/
_node.pklz
_inputs.pklz
desc-neutral_correlation.svg
_0x13eb83e355824a2708d409ea8dbfe4e2.json
result__atlas_corr_node0.pklz
desc-neutral_correlation.tsv
_report/
report.rst
_atlas_corr_node1/
_node.pklz
_0x2c551f03c95b45ac917f9caf672a88f4.json
_inputs.pklz
desc-congruent_correlation.svg
desc-congruent_correlation.tsv
result__atlas_corr_node1.pklz
_report/
report.rst
betaseries_wf/
dfb9084b801be401880a15ffb747f5a5a15543c2/
betaseries_node/
desc-neutral_betaseries.nii.gz
_node.pklz
_inputs.pklz
_0xf2e4cbdf258dc32033cb2d578dd1d722.json
result_betaseries_node.pklz
desc-congruent_betaseries.nii.gz
desc-incongruent_betaseries.nii.gz
_report/
report.rst
fmriprep/
dataset_description.json
sub-001/
func/
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-preproc_bold.nii.gz
sub-001_task-stroop_desc-confounds_regressors.tsv
sub-001_task-stroop_space-MNI152NLin2009cAsym_desc-brain_mask.nii.gz
sub-001/
anat/
sub-001_T1w.nii.gz
func/
sub-001_task-stroop_bold.nii.gz
sub-001_task-stroop_events.tsv
Collect results¶
corr_mat_path = os.path.join(out_dir, "nibetaseries", "sub-001", "func")
trial_types = ['congruent', 'incongruent', 'neutral']
filename_template = ('sub-001_task-stroop_space-MNI152NLin2009cAsym_'
'desc-{trial_type}_correlation.tsv')
pd_dict = {}
for trial_type in trial_types:
file_path = os.path.join(corr_mat_path, filename_template.format(trial_type=trial_type))
pd_dict[trial_type] = pd.read_csv(file_path, sep='\t', na_values="n/a", index_col=0)
# display example matrix
print(pd_dict[trial_type].head())
Out:
LH_Vis_1 LH_Vis_2 ... RH_Default_PCC_1 RH_Default_PCC_2
LH_Vis_1 NaN 0.090500 ... 0.104652 0.029874
LH_Vis_2 0.090500 NaN ... -0.110433 0.013504
LH_Vis_3 0.020199 0.213114 ... 0.220573 0.214643
LH_Vis_4 0.088036 -0.076339 ... -0.019350 -0.045271
LH_Vis_5 0.302583 0.375359 ... -0.249968 0.002706
[5 rows x 100 columns]
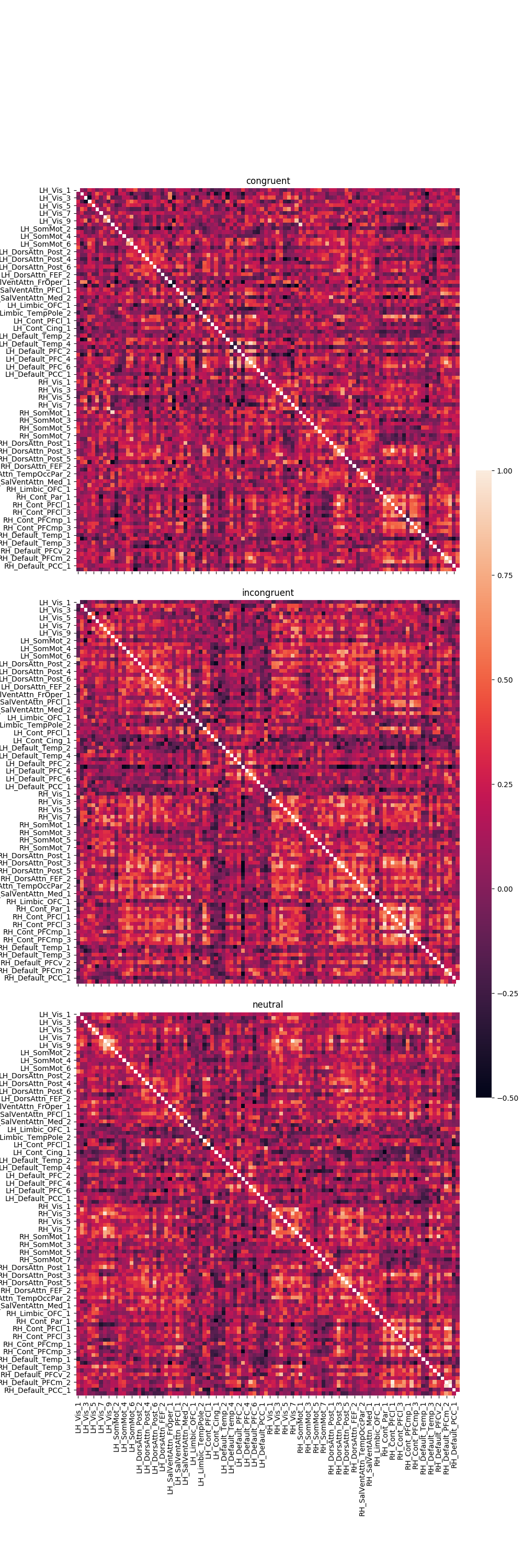
Graph the results¶
fig, axes = plt.subplots(nrows=3, ncols=1, sharex=True, sharey=True, figsize=(10, 30),
gridspec_kw={'wspace': 0.025, 'hspace': 0.075})
cbar_ax = fig.add_axes([.91, .3, .03, .4])
r = 0
for trial_type, df in pd_dict.items():
g = sns.heatmap(df, ax=axes[r], vmin=-.5, vmax=1., square=True,
cbar=True, cbar_ax=cbar_ax)
axes[r].set_title(trial_type)
# iterate over rows
r += 1
plt.tight_layout()
Out:
/home/docs/checkouts/readthedocs.org/user_builds/nibetaseries/envs/v0.4.0/lib/python3.7/site-packages/matplotlib/figure.py:2299: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.
warnings.warn("This figure includes Axes that are not compatible "
References¶
- notebook-1
Timothy D Verstynen. The organization and dynamics of corticostriatal pathways link the medial orbitofrontal cortex to future behavioral responses. Journal of Neurophysiology, 112(10):2457–2469, 2014. URL: https://doi.org/10.1152/jn.00221.2014, doi:10.1152/jn.00221.2014.
Total running time of the script: ( 1 minutes 59.376 seconds)